| DNA replication |
DNA (deoxyribose nucleic acid) is a biological macromolecule,which is the genetic data carrier: DNA is made of two complementary chains which forms a right.helix. Each chain is a sum of nuclieotides ; each nucleotide is compound of :
It exists four (4) bases of amino acids A (adenine), T (thymine), C (cytosine) and G (guanine). It's this complementarity whitch makes it possible the DNA to form a double helix. T joins A and C joins G During Mitosis, DNA will be replicated and thus there will be a doubling of DNA quantity. The purpose of this phenomenon is to distribute the same genetic information to all the cells of the organism. DNA replication is a semi conservative process, where each parental strand serves as matrix for synthesis of a new complementary daughter strand. There is a very important enzyme in this process, it's DNA polymerase (fig. 1).a Phosphate (able to establish the link between nucleotide) a Ribose (sugar) a base of amino acids (it's this element which differentiate the nucleotides)

It catalyzes the joining of desoxyribonucleoside triphosphate (dNTPs) to form a new DNA chain.A lot of other proteins and RNA specific sequences (called primers) are necessary. This system is compatible with a low frequency of errors in the replication mecanism.
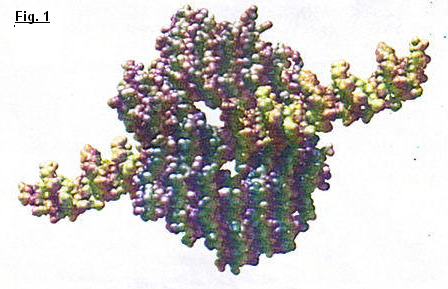
I. DNA polymerase :There are several types of DNA polymerase and each one have a specific function in DNA replication. DNA polymerase I, DNA polymerase II, DNA polymerase III in prokaryotic cells. This type was first identified in Escherichia coli in 1956 (nineteen fifty six). DNA polymerase alpha, beta, gama, delta and epsillon in eukaryotic cells. We know Gama DNA is responsible for replication of mitochondrial DNA and beta DNA is responsible for repairing of DNA damage. Others are most active in dividing cells. All DNA polymerases have two fundamental properties which allow a specific replication. (fig. 2) All the polymerase synthetize DNA only in the five primed to three primed direction. Adding dNTPs to the three primed hydroxyl group of a growing chain. The synthesis can take place only if a preformed primer strand is present. It's the main difference between DNA and RNA, because RNA can initiate the synthesis of a new strand in absence of a primer. However it's this property which confer the high fidelity of DNA replication that is required for cell reproduction.
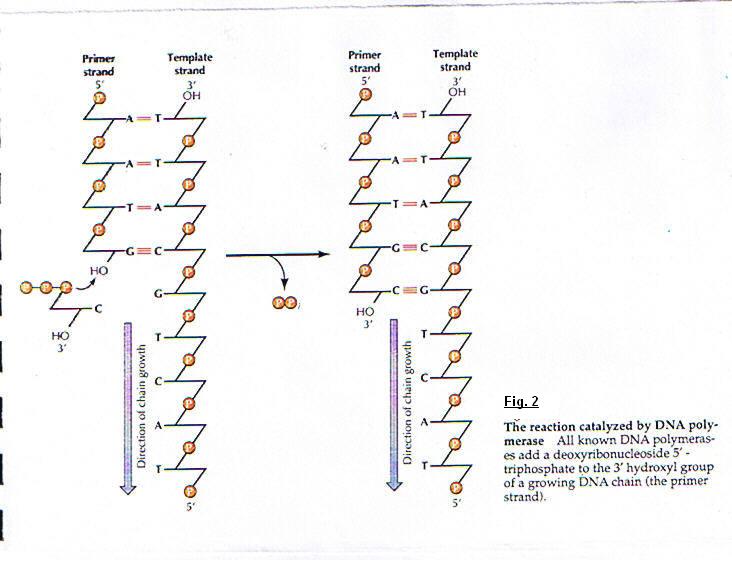
The small pieces are called Okasaki fragments (the fragments have the name of discoverer) ; and joined by the action of DNA ligase. (fig. 3) The short fragments of RNA serve as primers for Okasaki fragments. The enzyme which synthesizes short fragments of RNA (three to ten nucleotides long) is called primase. (fig. 4) To form a continuous lagging strand of DNA : the RNA primers must eventually be removed from the Okasaki fragments and replace with DNA. In Escherichia coli primers are removed by the combined action of RNase H, and an enzyme which can degrades the RNA strand when it's in hybrid form (DNA-RNA), this enzyme is polymerase I.
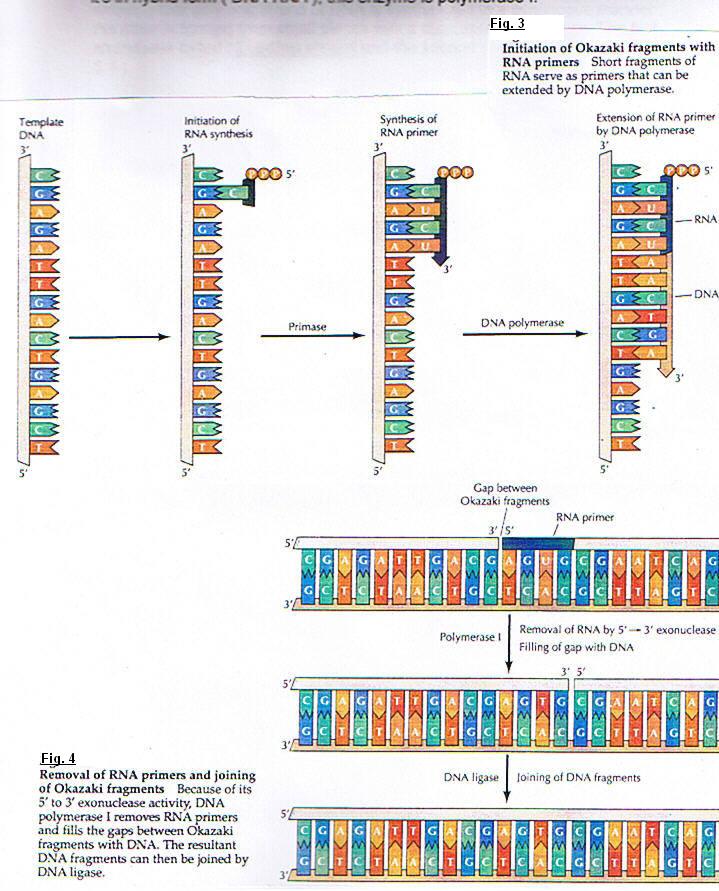
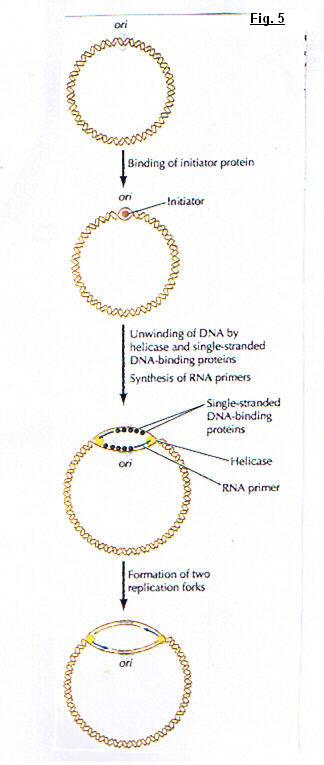
(fig. 5)DNA molecules contain two replication forks. At this place the parental strands of DNA are separated by the rupture of a hydrogen-bond. This opening constitute replication eye and two new daughter strands are synthetized. Each fork run in opposite direction. so, under these conditions, the synthesis poses an important problem; because we showed the DNA polymerase catalyses the polymerisation only in five primed to three primed and we know that the two DNA strands are antiparallel. Several experiments showing that only one strand of DNA is synthesized in a continuous manner ; the other is formed from small pieces with a discontinuous process. The first strand was called : Leading strand and the second : Lagging strand. (fig. 6)
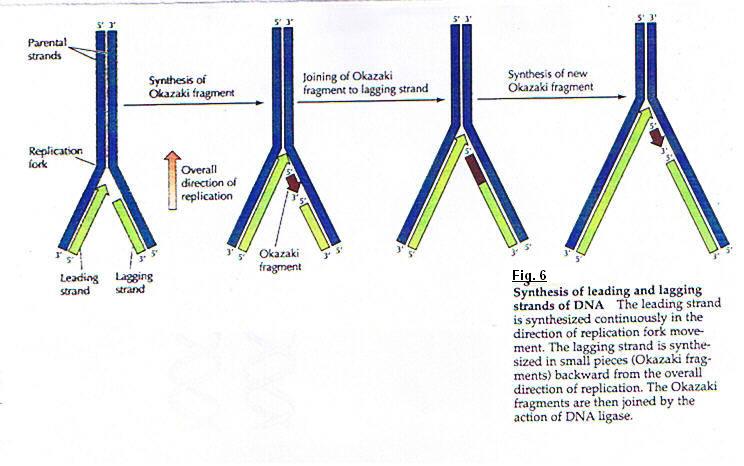
The polymerase I acts as an exonulease that can hydrolyze DNA (or RNA) in either the three primed to five primed or five primed to three primed direction. The action of polymerase I as a five primed to three primed exonuclease removes ribonucleotides from the five primed ends of Okasaki fragments and replace them with desoxyribonuleotides. These DNA fragments are joined by DNA ligase.
Others proteins unwind the template DNA and stabilize the single strand regions.
(fig. 7) Helicases are enzymes that catalyze the unwinding of parental DNA, coupled to the hydrolis of ATP. DNA can be copied only on single strand state. When helicase unwinds parental DNA, the DNA ahead of the replication fork is forced to rotate. (fig. 8) This rotation can block replication. This problem is solved by Topoisomerases : enzyme that catalyze the reversible breakage and rejoining of DNA strands.
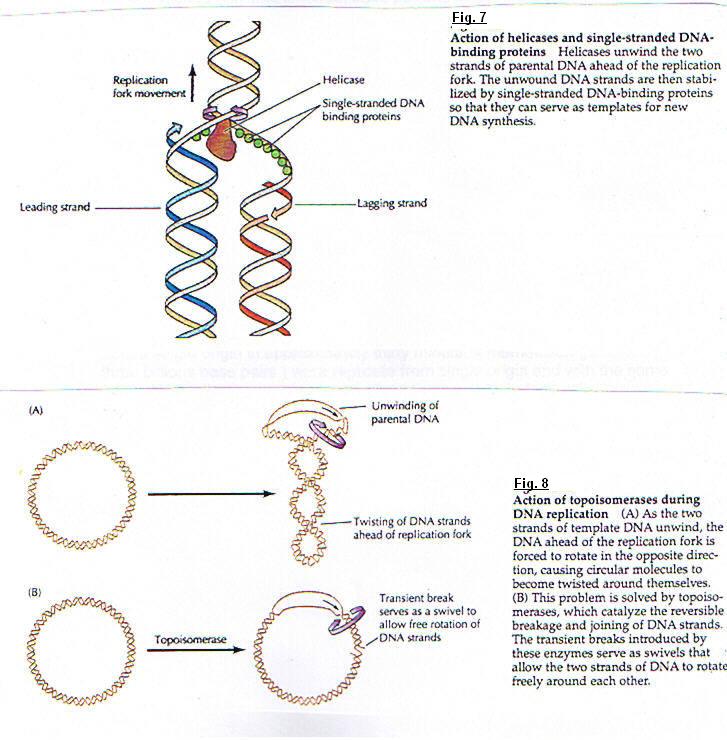
The enzyme involved in DNA replication act in a coordinated manner to synthesize both leading and lagging strands of DNA simultaneously at the replication fork. DNA polymerase must accomplish the formation of dimers. Each monomer syntetizes one strand. The template lagging strand is thought to form a loop at the replication fork so that the polymerase subunit engaged in lagging strand synthesis moves in the same overall direction as the other subunit, which is synthesizing the leading strand.
III. The fidelity of replication :The accuracy of DNA replicaition is critical to cell reproduction, and estimates of mutation rates indicate that the frequency of error during replication corresponds to only one incorrect base per one billion nucleotides incorporated.
A major mechanism reponsible for the accuracy of DNA replication is the proofreading activity of DNA polymerase. It's the five primed to three primed as well as three primed to five primed exonuclease activity which can participates in proofreading newly synthesized DNA
IV. Initiation of replicationThe replication of both prokaryotic and eukaryotic DNA starts from a unique sequence called origin of replication, it's a specific binding site for proteins that initiate the replication process. The key step is the binding of an initiator protein to specific DNA sequence. This initiator protein begins to unwind the origin DNA and helicase and single strand DNA binding proteins then act to continue unwinding and exposing the template DNA ; and primase initiates the synthesis of leading strand. Two replication forks are formed and move in opposite direction.
Escherichia coli has a genome of four millions base pairs and is replicated from a single origin in approximately thirty minuts. If mammalian genomes (three billions base pairs) were replicate from single origin and with the same rate, DNA replication would require about three weeks.In fact, cells mammalian genomes are typiccaly replicated within a few hours ; necessitating thousands of replication origins (thirty thousands in human genom).